Expected progeny differences (EPDs) are a widely utilized tool in making genetic decisions centered around breeding objectives. With the progression of DNA technology, the industry has been able to incorporate genomics into the numbers that are used to calculate EPDs. Prior to inclusion of genomic information, we relied on pedigree-based relationships that operate on the averages. This assumes a 50% contribution from the dam and 50% contribution from the sire to the breeding value of the offspring. However, an animal’s actual genetic merit or it’s EPD value for a given trait can be greater or less than that of its parents, depending upon how the numerous random combinations of genetic material contributed by the sire and dam come together and are expressed in the offspring.
The lacking component of historical genetic assumptions is accuracy. As a bull sires more offspring and those records were reported the EPDs would change to reflect offspring performance and accuracy would increase. By including genomic information in EPD calculations, it increases the accuracy of younger sires. For EPDs with large amounts of records such as birthweight, genomics can be the equivalent of already having 25 offspring records submitted on that animal prior to ever breeding a cow.
Sampling methods to collect tissues for identifying the genetic makeup of the animal can include blood samples, hair samples, or tissue samples such as ear punches. Through the collection of DNA samples from a calf and its parents, we can make better estimations as to how an animal will relate to each parent, or even grandparent, on a genotype level. This understanding gives more accuracy to knowing a young animals EPDs which gives greater precision to predict the impact on performance of future offspring.
Table 1 is a comparison of full sibling bull (twin) calves that represent genetics out of the same sire and dam that have been DNA tested and that information utilized in the calculation of their EPDs.
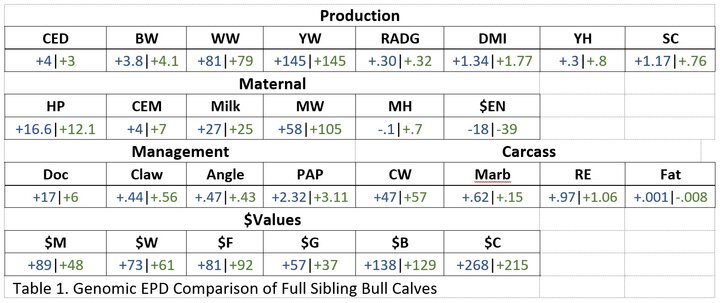
In this comparison, we can see these bulls are similar in CED, WW, YW, and Milk for some examples. In contrast, we can see where these bulls differ in SC, MW, and Doc. The differences and similarities are echoed through the comparisons of the bulls’ $Values. Calf #1 displays opportunities to be the better selection when looking to retain females ($M). On the other hand, if retaining ownership of calves through the finishing period and marketing as a live animal, Calf #2 would produce offspring with higher profit potential.
Genomic-enhanced EPDs are the best estimate of an animal’s genetic value as a parent, combining all available sources of information. This permits higher accuracies for EPDs on younger animals. With the inclusion of genomics, we can better understand how an offspring relates to its parents from trait to trait, which creates an opening for increased certainty that mating selections will progress towards breeding objectives more rapidly and as intended.
Interviews with the authors of BeefWatch newsletter articles become available throughout the month of publication and are accessible at https://go.unl.edu/podcast.

